Here are some stem cell stories that caught our eye this past week. Some are groundbreaking science, others are of personal interest to us, and still others are just fun.
Two hearts beat as one, or not
Sorry for the pre-Valentine’s Day buzzkill but stem cell research published this past week points to a very unromantic discovery: two hearts do not beat as one. The study, out of Rockefeller University, and published in the Journal of Cell Biology, sought to understand the limited success of clinical trials in which stem cell-derived heart muscle cells, or cardiomyocytes, are transplanted into the heart to help repair tissue scarred by disease or a heart attack.
If you’re a regular at The Stem Cellar, you’ll recall that just last Friday we summarized published experiments that suggest the cardiomyocytes used in successful trials do not grow new tissue themselves but instead heal the heart indirectly by releasing proteins that stimulate repair.
The research team behind this week’s study instead reasoned that the transplanted cardiomyocytes do indeed integrate into the heart tissue, but they fail to contract properly with the undamaged heart cells. So, the thinking goes, the transplanted cells do nothing to restore the heart’s ability to beat at full strength.

A two-cell “microtissue” contains a mouse embryonic stem cell-derived cardiomyocyte and a mouse neonatal cardiomyocyte. The lower panel shows the traction forces generated as the two cells contract; the stronger, neonatal cardiomyocyte produces more force than the weaker, stem cell-derived cardiomyocyte. Credit: Aratyn-Schause, Y. et al. J Cell Biol. 2016 Watch video here: http://medicalxpress.com/news/2016-02-muscles-on-a-chip-insight-cardiac-stem.html
To test this hypothesis, the researchers devised a two-cell micro-tissue made up of a single mouse cardiomyocyte and a single cardiomyocyte derived from either mouse embryonic stem cells or induced pluripotent stem cells (iPS). This “muscle-on-a-chip” showed that the two cells are able to physically connect up and even beat in sync with each other. But, the embryonic and iPS-derived cardiomyocytes beat less strongly than the native cell. Based on computer simulations, this imbalance made the micro-tissue beat less efficiently. A university press release picked up by Newswise includes a short yet fascinating video of the differing strengths of the beating heart cells (click on image above).
With this micro-tissue in hand, the team aims to find a way to fix this imbalance, which hopefully would make cell therapies for heart disease more potent.
Your Own Private Micro-liver
Enough about micro-hearts, let’s talk micro-livers.
In a report published on Monday in PNAS, a multidisciplinary UCSD team of engineers and biomedical researchers described the creation of a bioprinted 3D liver model made from human iPS-derived liver cells, or hepatocytes. The hepatocytes are imprinted on a surface in hexagonal shapes, the kind seen in the complex microarchitecture of the human liver. These structures were also seeded with two other cell types: endothelial cells, which form blood vessels, and fat cells, which support the health of hepatocytes. Including these relevant cell types in the “micro-liver” design resulted in a 3D cell culture that not only mimics structures but also replicates functions found in a natural liver.
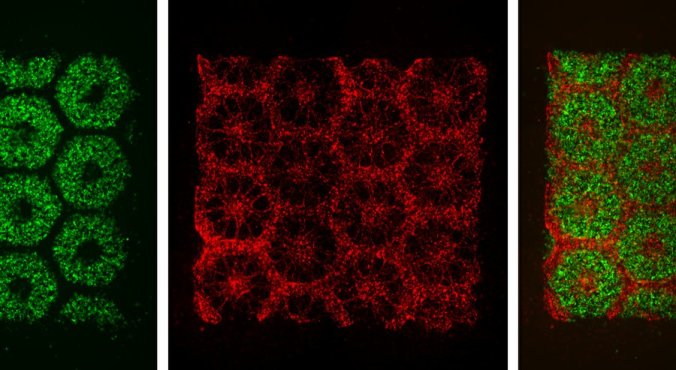
The 3-D-printed parts of the biomimetic liver tissue include: liver cells derived from human induced pluripotent stem cells (left), endothelial and mesenchymal supporing cells (center), and the resulting organized combination of multiple cell types (right). — Chen Laboratory, UC San Diego
This is a really exciting development for improving drug safety. A big concern of any new drug coming on the market is its potential liver toxicity, formally known as DILI (drug induced liver injury), the most common cause of liver failure in the U.S. Although animal studies and clinical trials carefully test for the potential of DILI, that doesn’t guarantee the drug will be safe in all individuals. And because this liver model was designed using human iPS cells – which can be derived from anyone with a simple skin biopsy – it has the potential to serve as a personalized drug screening device as well as a disease-in-a-dish model for studying inherited forms of liver disease.
As Bradley Fikes, San Diego Union Tribune’s biotechnology writer, mentions in an excellent summary of the publication, beyond drug screening and disease-in-dish modeling, this bioprinting process could also one day make it possible for researchers to reach the “holy grail” of tissue engineering: building an entire organ.
Finally! The Bloody Holy Grail
While that holy grail remains on the horizon, Stanford researchers are nearly holding the goblet in their hands. Based on a Nature report published yesterday, a team led by CIRM grantee Irv Weissman have found a long sought after cellular tag that can fish out a very specific type of hematopoietic stem cell (HSC), or blood-forming stem cell, from bone marrow.
Almost thirty years ago, Weissman identified HSCs, which have the ability to form all the cell types of the blood. Since that time, scientists have struggled with fully understanding how HSCs are maintained in the body and, in turn, how to grow them in the laboratory.
The source of this problem is due to the fact that most HSCs are so-called short term HSCs because they eventually lose their “stemness”; that is, their ability to divide indefinitely. Only a small fraction of HSCs are of the long-term variety. To really understand how the body sustains a life-long supply of HSCs, it’s necessary to have a method to pick out just the long term HSCs.
So scientists in Weissman’s lab set out to do just that. Starting with a list of 100 genes that are known to be active in the bone marrow, they looked for genes that are turned on only in long term HSCs. After a painstaking, systematic method that took two years, the team narrowed down the list to just one gene that was unique to long term HSCs.
Co-lead author James Y. Chen, a MD/PHD candidate at Stanford, described the significance of this effort in a university press release:
James Y. Chen
“For nearly 30 years, people have been trying to grow HSCs outside the body and have not been able to do it — it’s arguably the ‘holy grail’ in this field. Now that we have an anchor, a way to look at long-term HSCs, we can look at the cells around them to understand and, ideally, recreate the niche.”
Older Dads and The Selfish Sperm
We wrap up the week with a PNAS publication that got a wide range of coverage by the likes of BBC News, Gizmodo and Cosmos in addition to the usual suspects like Health Canal. Not too surprising given the topic including selfish sperm and chopped up testicles.
Research over the past decade or so has made it increasingly clear that biological clocks not only tick for would-be moms but also dads. At first glance, it makes sense: older fathers have had more time to accumulate random DNA mutations in their spermatogonia, the stem cells that produce sperm. But studies of Apert syndrome, a rare disease causing defects in the skull, fingers and toes, has put this hypothesis in question.
Back in 2003, a research team at Oxford University found the mutation in spermatogonia that causes Apert syndrome occurs 100 to 1000 times more frequently than would be expected if it were merely due to a random mutation (the Apert syndrome is not inherited because males with the disease rarely go on to have children).
So what’s going on? To answer that question the Oxford scientists collaborated with a USC research team who (men: you may not want to read the rest of this sentence, this is your only warning) chopped up human testicles – ones that had been removed for unrelated medical reasons and donated – in order to reconstruct a three-dimensional map of where these Apert syndrome mutations were occurring. If the mutations were merely random, the affected spermatogonia would have been evenly distributed throughout the testicle. Instead, the team found clusters of cells carrying the mutation.
This results confirms a “selfish sperm” hypothesis in which the mutation provides a selective advantage to the affected sperm cells allowing them to out compete other nearby sperm cells, much like a cancer cell that multiples and gradually forms a tumor. The study serves as more sobering news to otherwise healthy older dads that they may have a higher risk of passing on harmful mutations to their offspring.
Like I said, sorry for the buzzkill. Happy Valentine’s Day weekend!
